This post dives into the sklearn decision tree building process.
The node_split method within the BestSplitter class is probably the most element implementation that I will say within the whole decision tree module. Not only it entails the implementation of the splitting process, but also demonstrated a myriad of core computer science concepts. Now let’s go through these 200 LOC and enjoy.

In the method documentation string, it is mean to “Find the best split on node samples[start:end]”. Variable samples store all the samples, or each node_split is not necessarily running against the whole samples at every split. During each split, its range will target at the start:end range which will find the rotation later.
Meanwhile, at each node_split, its inputs include not only the latest impurity process, it also contains a pointer split to the split record, last but not least, for efficiency purpose, the splitters keep track of the number of constant_features so we can speed up the splitting by avoiding constant features, hence the comments below:
# Sample up to max_features without replacement using a# Fisher-Yates-based algorithm (using the local variables `f_i` and# `f_j` to compute a permutation of the `features` array).## Skip the CPU intensive evaluation of the impurity criterion for# features that were already detected as constant (hence not suitable# for good splitting) by ancestor nodes and save the information on# newly discovered constant features to spare computation on descendant# nodes.
First, let’s look at the outermost loop and its condition:
- f_i > n_total_constants
- n_visited_features < max_features
- n_visited_features <= n_found_constants + n_drawn_constants
condition 1 and (condition 2 or condition 3)
max_features is a an input variable that can be changed by the user, but default to be the total number of features. When float, it will analyze pick a fraction of all the features during the split or hard coded to be a specific number of features.
The implementation takes into account constant variables, and has several flags/counter to keep track in order to improve the performance. However, that might not help the reader so we will assume all the columns are not constant so any counter related to constant will be zero.
In that case, the while loop condition will be (f_i > 0 and n_visited_features < max).
n_visited_features is initialized to be zero and increases by 1 at each loop.
f_i was initialized to be n_features outside the loop, and at each loop f_j is randomly drawn from (n_drawn_constant, f_i – n_found_constants) ~ (0, f_i) when there is no constants. Of course, if we do have constants, f_j is meant to randomly select a feature that is not drawn nor not found constant.
When you draw the feature, the code checks what kind of feature this is, and update the related counter accordingly.
For example, as it is random sample without replacement, if it has been drawn, then it should not be drawn again, hence, f_j has to a random integer greater than the n_drawn_constants. f_j > n_drawn_constant.
At the same time, if f_j is smaller than n_known_constants, which means that we might have already known it is a constant without drawn yet (cache), then, we will mark this constant column as a drawn column by switch columns, append to the end of the drawn_constants range and update the n_drawn_constant counter by one.
Of course, we don’t know yet, we will update f_j by adding the identified/found constants so f_j is now pointing to the f_j th unidentified column and use it as the current feature.
Once we select a feature, then we will copy the column values to a new array Xf and sort the values. After we sort the values, then we will check to see if the column is close enough to be constant by comparing the min Xf[start] and max Xf[end-1] and see if the difference is smaller than feature_threshold. If so, of course, we will update the found_constants by 1 and total_constant by 1.
Of course, after all, if is not known constants, or if it is new but not constant, then we will go and find the split points by decreasing f_i by 1 and switch values of feature f_i and f_j. In that case, our randomly selected features will always be at the end of the list and slowly building up to the start of the queue.
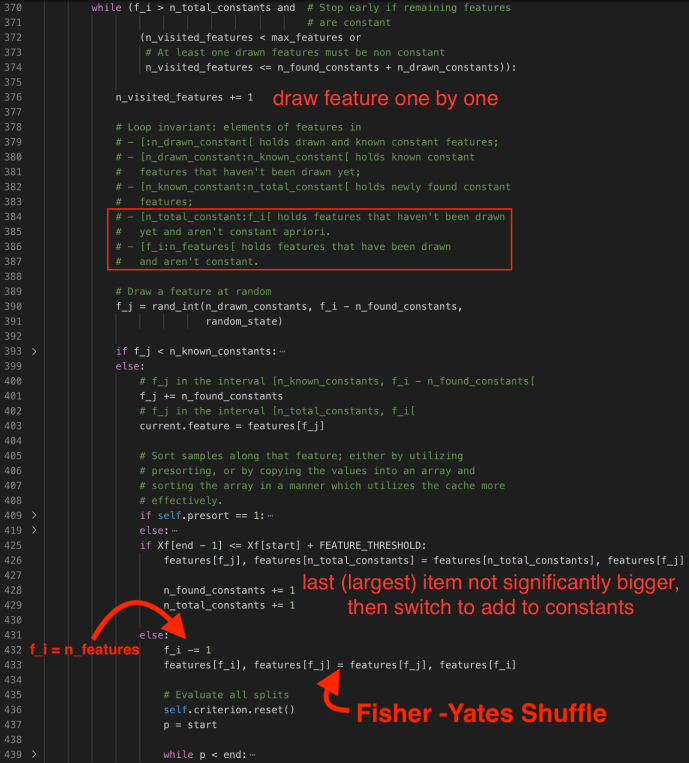

When the feature is properly sampled, they first initialize the position – variable p to be the start, the record index of the first sample. And it will go through each record. In addition, when it moves on to the next record, they FEATURE_THRESHOLD to make sure that it was not numerically equal, if so, they will skip to the next record which is materially different.
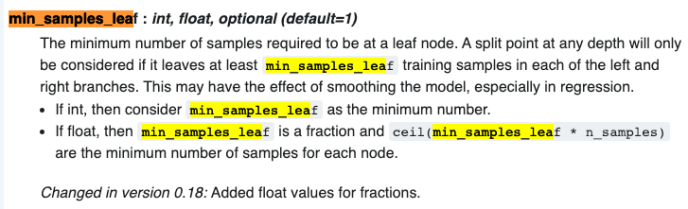
When sklearn is trying to find a split point during a continuous range, there is a very important user input called min_samples_leaf, which is defaulted to 1. When you build a decision tree, theoretically, you can train your model to be 100% accuracy during the training stage by creating a tree path which each unique record falls into its own path. Your model will for sure be big, and at the same time, you might be cursed with overfitting your data that during the prediction stage, the accuracy will be drastically lower. To avoid that, min_samples_leaf can be used to make sure you only create split point when the children has more than min_samples_leaf data. Here is the official documentation.

The same idea applies to the min_weight_leaf in which instead of samples count, the requirements hold true for sum of weight.
After these two checks, they will calculate the criterion improvement and calculate with the record keeper until they loop through all positions and find the best split point and save that position as the best.

After they find the best split point, they will “Reorganize” and actually split the samples using the best split point. Keep in mind that X still holds the raw samples and are NOT sorted. while p:= start < partition_end := end is an interesting way of splitting the data and sort into two subgroups in which left part of the array are always smaller and the right is always larger.
Here is a small Python script to demonstrate how it works, just to clarify, the data is still not completely sorted.


After the “Reorganization”, the memory of constant features will be copied and return the best split point.
In summary, in this post, we have covered the split_node method with in the BestSplitter class. We looked at how they use Fisher_Yates random sample the features, for a given feature, how it find the best splitting point and at last, how it cleaned everything up and pass on to the children.